GeneBench and current agentic capabilities in the life sciences
A new benchmark for evaluating real scientific workflows in genomics and quantitative biology
tl;dr Most benchmarks don't capture the complexity and open-ended nature of actual scientific analysis. We introduce GeneBench, a new benchmark that attempts to begin measuring agentic capabilities at achieving this kind of analysis in genomics and quantitative biology. In the current 103-evaluation suite, GPT-5.5 Pro reaches 33.2% overall, while GPT-5.5 reaches 25.0% at xhigh reasoning effort. Current models struggle with iterating on initial hypotheses and allowing intermediate findings to alter an analysis trajectory, but they are rapidly improving.
Introduction
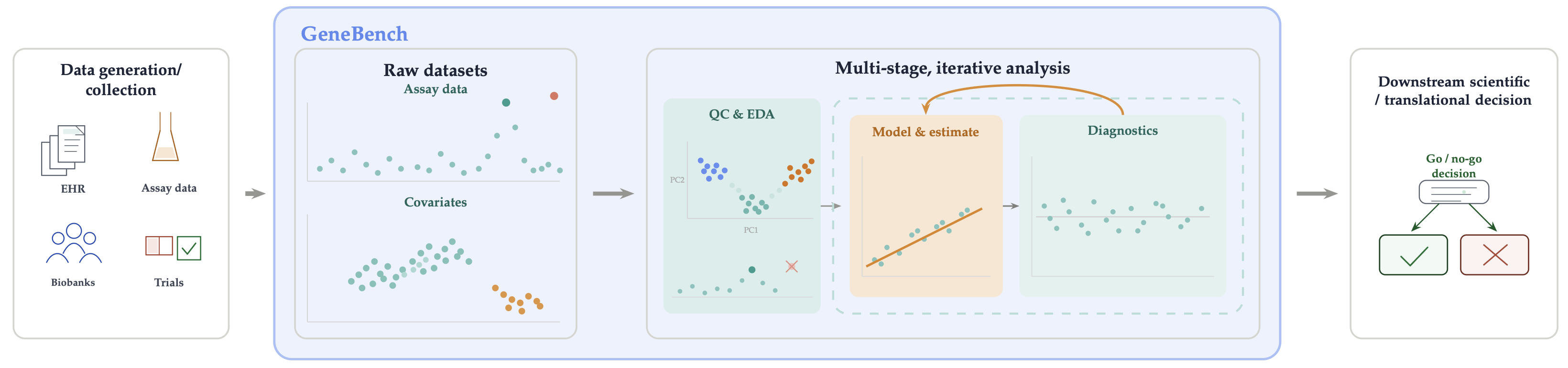
In this post we introduce GeneBench, a new benchmark developed in collaboration with OpenAI that evaluates agentic AI systems on their ability to execute realistic, multi-stage scientific analyses requiring navigation of ambiguities ubiquitous in scientific research. GeneBench is focused on problems in genomics and quantitative biology, but the underlying analytical pattern is common to end-to-end scientific analysis workflows across the computational life sciences.
The full preprint can be found here.
In this post I provide more context around the original motivation for constructing this benchmark, briefly discuss the high-level results, and close with a few thoughts on where things might be going next.
Brief background
I have been working in statistical genetics for the last ~decade across startups as both a computational researcher in an IC capacity as well as a research team lead working on methods development and applied research. My day job is as a research engineer at Herasight, a startup offering best-in-class polygenic embryo screening. Check them out if you're considering having kids soon.
Motivation and scientific research in the real world
Agentic systems such as Codex and Claude Code have taken engineering and technical fields by storm over the last six months. I have been using them in my day job as a researcher in genomics since Claude Code was initially released, and my rough sense is that the total factor multiplier on my own productivity has probably been ~5-10x, with the value steadily increasing over the last year. For example, I have been able to develop and ship sophisticated MCMC algorithms with hardware-accelerated kernels in weeks that would likely otherwise have taken me at least a year to come up with and implement myself. Like many others, I haven't written a line of code in months.
Nevertheless the main axis of improvement over the last year has largely been on the quality of execution rather than on the level of delegation I find myself comfortable with giving agents. They are more than good enough to execute a sophisticated statistical, bioinformatic, or scientific workflow as long as I tell them "apply this method to this data and tell me what you find," but I still don't trust them to execute an experiment end to end in the sense of "here's some data, here's my hypothesis, go and check if it's right." I have to intervene fairly often to course-correct, redirect, and provide higher-level guidance.
This is in part because scientific research inherently contains a great deal of ambiguity in what constitutes the "right approach" and where there may be no single "correct" answer. For example, (1) multiple defensible approaches might exist, (2) results may hinge on researcher degrees of freedom, (3) one's initial hypothesis may be totally incorrect for reasons that can only be discovered during the process of analysis, and (4) the data one works with may be broken in some fundamental but non-obvious way.
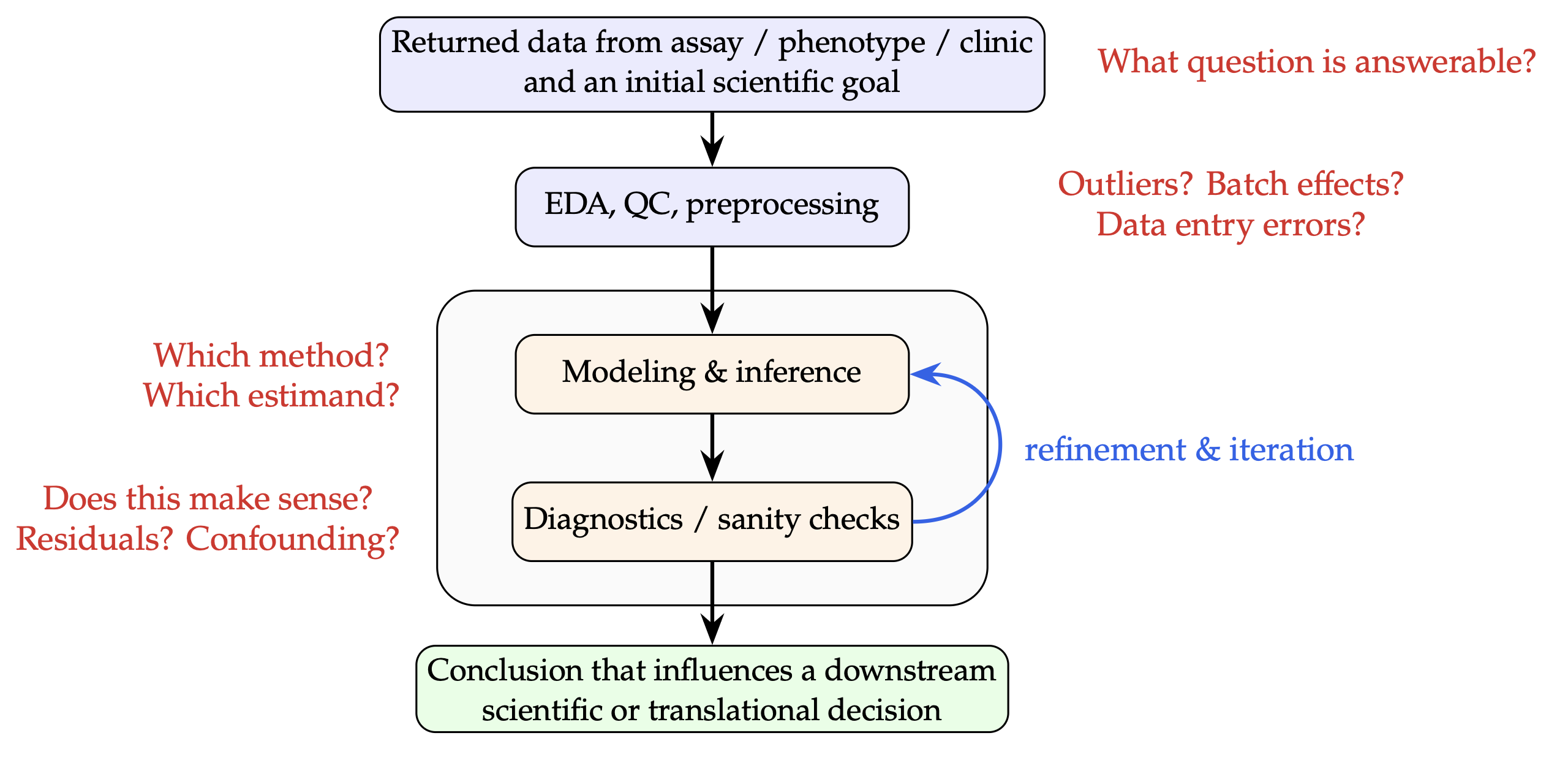
Figure 2 shows a high-level sketch of a typical analysis in real-world biology research, where one is presented with one or more datasets from any number of data collection pipelines (e.g. biological assays, biobanks, electronic health records, etc.) and where the goal is to arrive at a conclusion which informs a downstream decision (whether business, clinical, or otherwise). Successfully navigating this graph demands the ability to make a sequence of high-fidelity judgment calls as part of an iterative process of including backtracking, iteration, and exploration.
Existing benchmarks for AI capabilities in the life sciences do not adequately capture this aspect of research
The existing literature on AI capabilities in biology generally falls into one of two buckets. The first consists of benchmark-style studies such as GeneTuring, LAB-Bench, LABBench2, BixBench, BAISBench, SpatialBench, scBench, and CompBioBench, where agents are given tightly scoped problems with validated answers and graded on whether they recover the correct result. The second consists of higher-level "AI scientist" studies like AI co-scientist, the Virtual Lab, and CellVoyager, where the question is whether a system can produce apparently novel hypotheses, discoveries, or accelerate research.
Personally, my time these days is not spent on executing the tightly scoped, rote analyses that the first bucket evaluates (the models can already execute those reliably enough), and the systems described in the second bucket require bespoke setup and infrastructure and are aimed at highly specific and narrowly scoped optimization tasks. What is largely missing is evaluation of the ordinary, generic "unit" of scientific work between these two extremes: recovering a valid analysis from messy real-world data under limited guidance. In my view, this is where current agentic systems have the greatest potential to accelerate everyday scientific research.
GeneBench
Overview
GeneBench is a new benchmark focused on problems in genomics and quantitative biology that aims to partially fill this gap. Each problem in GeneBench is a self-contained analysis problem where an agent is given raw, messy datasets intended to reflect what a scientist would receive in practice, along with a "minimum viable prompt" which describes the setup and a target quantity to estimate, but does not provide any further guidance. The scope of each problem is intended to reflect an analysis that would take a senior scientist anywhere from 10-40 hours to execute without the help of AI.
Each problem is designed around obstacles common in real scientific work but underrepresented in existing benchmarks: messy data, measurement error, selection bias, confounding, QC failures, and the need to choose among competing model classes. Problems are constructed with multiple dependent decision points: substantive inferential forks where a plausible wrong choice changes the downstream analysis, such that errors propagate through the chain and into the final graded target.
Each problem uses simulated data for which the full data-generating process is known. This allows us to (1) deliberately build in ambiguities and data/statistical issues reflective of real-world problems, (2) tune and calibrate difficulty of each decision point in the inferential chain, and (3) ensure that the graded answer results from the single path through the garden of forking paths of data/QC/modeling decisions that remains viable when all of the data are taken into account.
This last point is particularly important, because the open-ended nature of scientific research is also precisely what makes it difficult to construct and grade such problems in an objective manner. Suppose you are trying to evaluate an agent's ability to detect and trim outliers from some distribution of a QC metric. In real data, these distributions are usually not glaringly bimodal with a 'good' and 'bad' mode which makes it obvious where to draw the threshold; often the choice comes down to some heuristic the researcher chooses to apply, and a threshold of z=1 may well be as defensible from a scientific standpoint as z=2. If an agent is graded on a quantity that depends on where it decided to threshold, what then should be considered the "correct" answer --- the result from trimming at z=1 or z=2? One could consider both values correct, but most real-world analyses require chaining together multiple such decisions, and it then becomes unclear how best to deal with this combinatorial explosion in the context of evaluation.
Thus, for each decision point in each GeneBench problem, we require that there be an aspect of the data that points clearly to one "correct" approach -- for example, in cases where QC thresholding is a decision point, we deliberately make it obvious where the threshold should be drawn, and make the final graded answer insensitive to the exact choice of threshold, as long as some thresholding is performed.
Without full control over the data generating process, achieving this property for problems would be significantly more challenging.
Scope
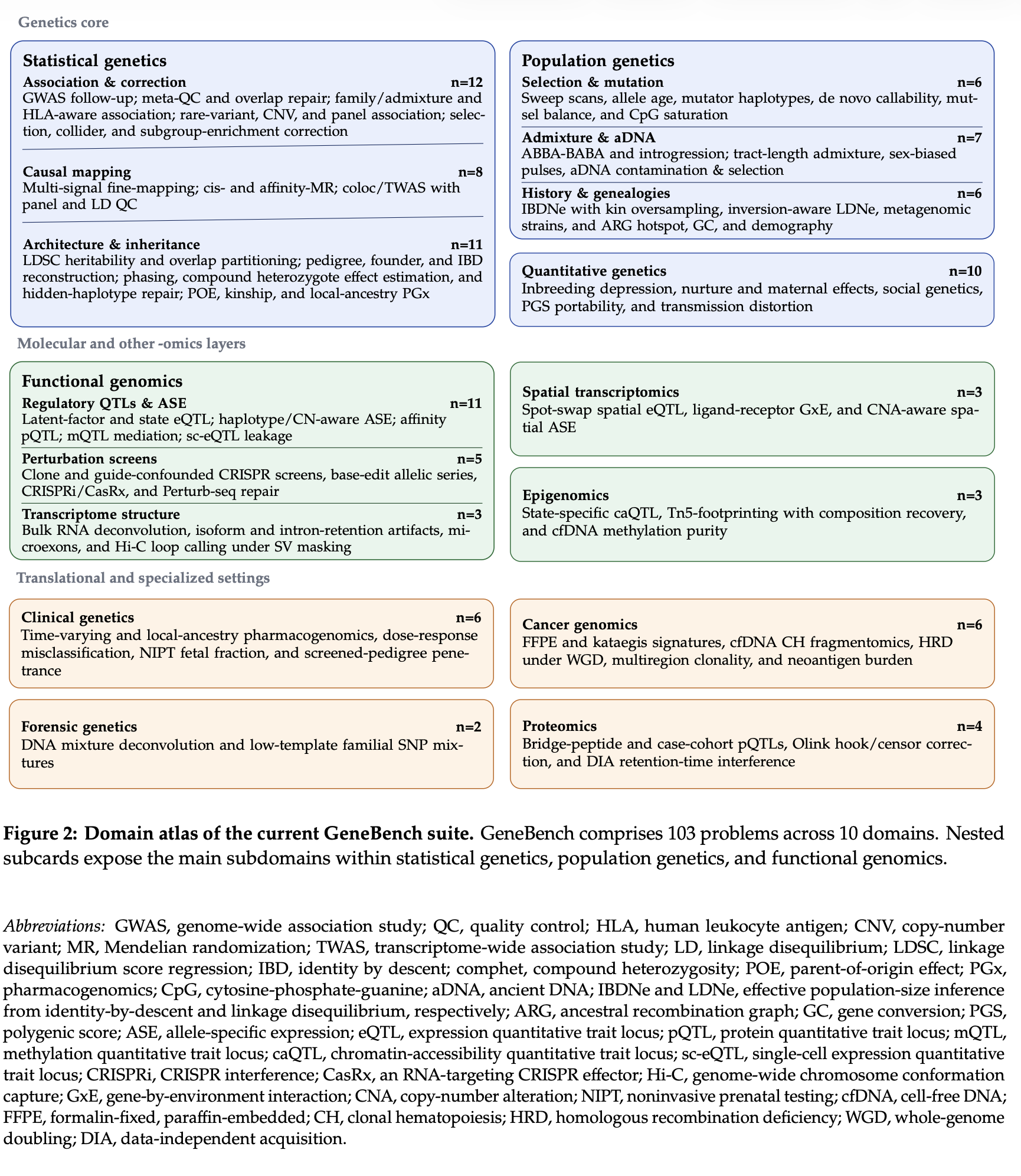
The problems in GeneBench are designed to reflect a fairly wide range of questions that scientists in both industry and academia actually spend time on. About two thirds of the suite covers topics in statistical, population, and quantitative genetics. These include industry-facing questions like whether a GWAS signal survives follow-up strongly enough to justify advancing a target, and which gene or protein at a locus should actually be prioritized for follow-up. They also include more academic questions like whether an observed pattern is better explained by selection or demography, or what pedigree, haplotype, or ancestry reconstruction is actually supported by noisy data. The remainder of the questions cover functional genomics and adjacent 'omics and quantitative biology settings, including spatial transcriptomics, cancer genomics, proteomics, clinical genetics, forensic genetics, and epigenomics.
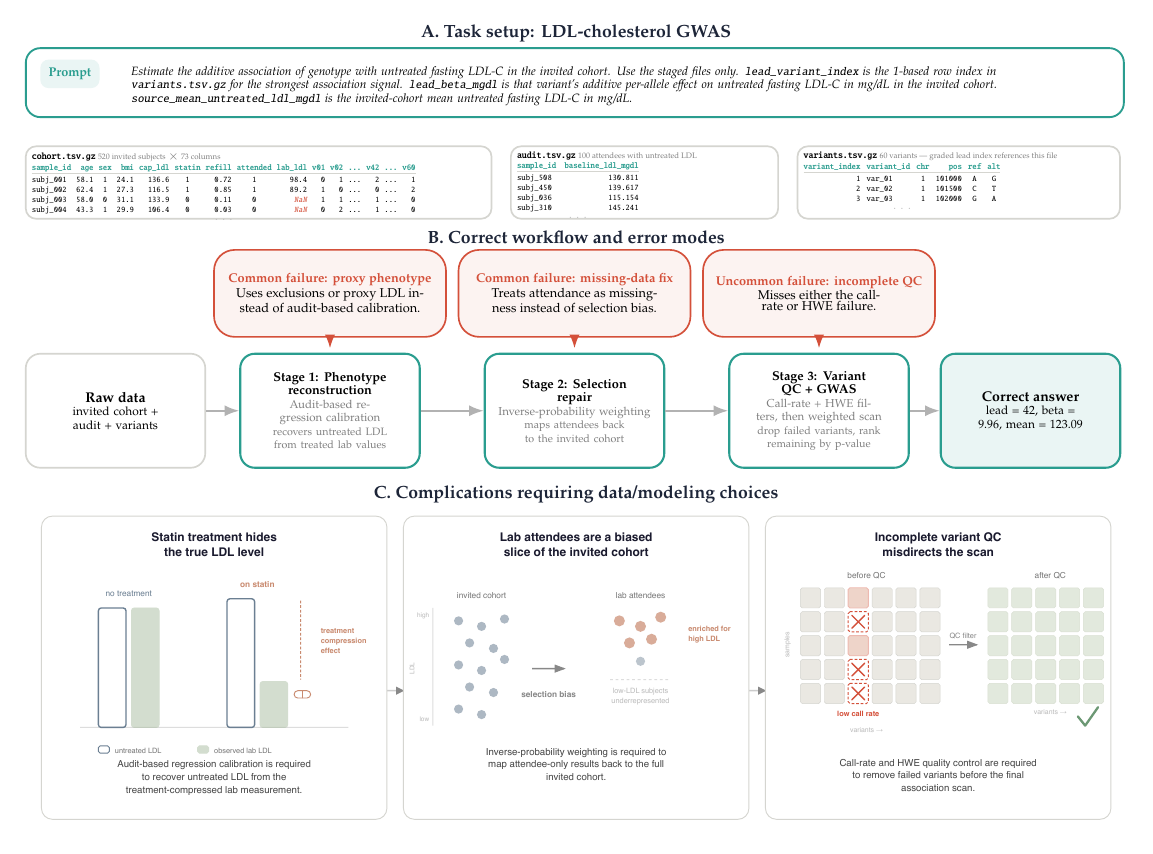
Further details of the benchmark and some representative problems as well as a worked example of a GeneBench problem can be found in the paper or the OpenAI blog post.
Agent environment
The environment agents are provided with is deliberately minimal, and is just a Linux environment in a Docker container with Python and standard scientific computing libraries.
Each problem comprises a set of data files and a "minimum viable prompt" that defines the target quantity as well as a brief description of the file contents. Agents are instructed to return a json with defined key-value pairs containing their final estimates of the target quantity, which are then graded against the ground truth. Currently, grading is done in a binary, pass/fail manner.
Results
Overall performance
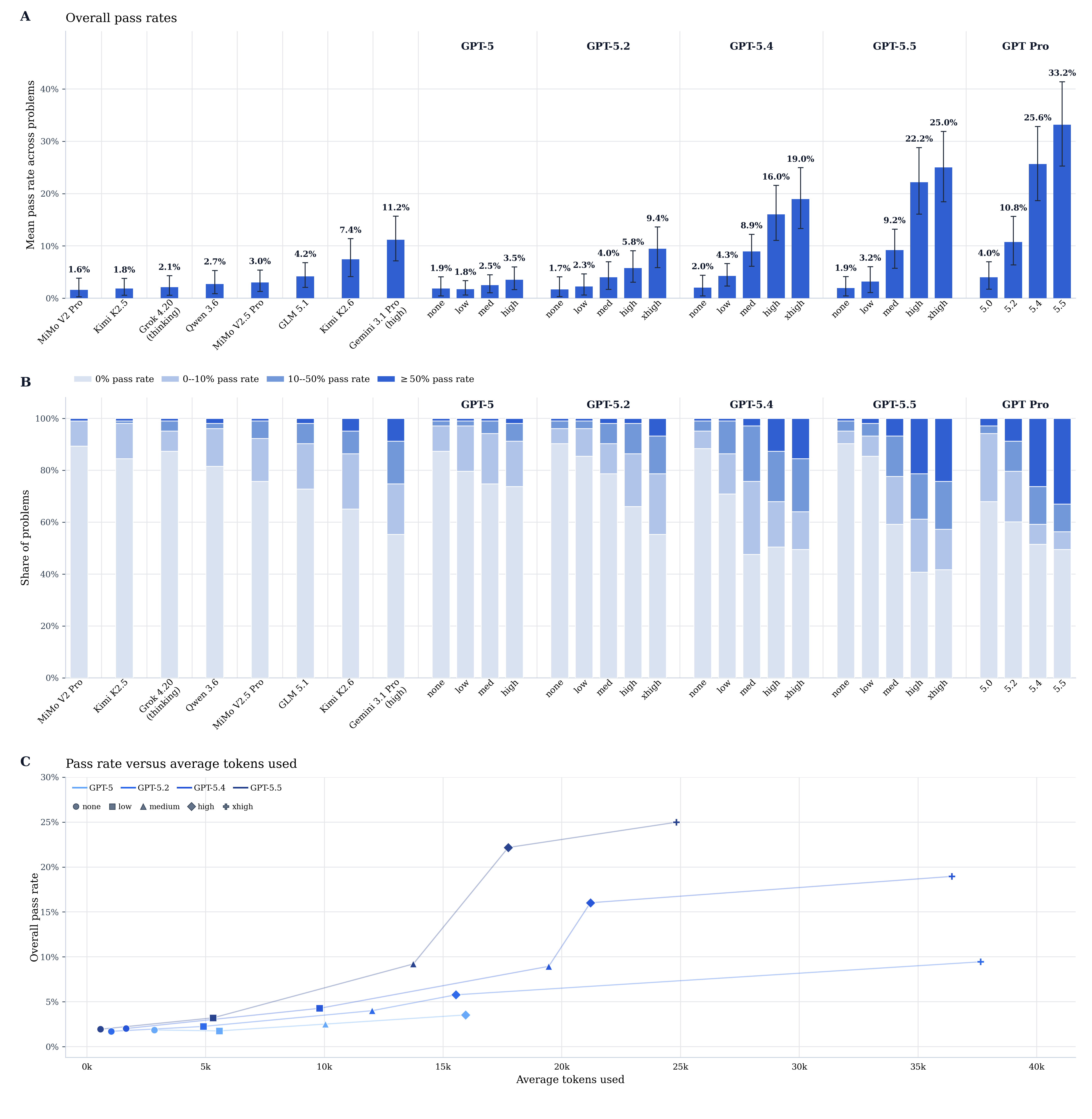
The headline result is that current frontier models perform moderately well on the existing problems, with GPT-5.5 Pro achieving the highest overall pass rate at 33.2%, and the strongest non-OpenAI model, Gemini 3.1 Pro, achieving 11.2%.
The Pro runs are reported separately from the mainline GPT effort-scaling curve; on that curve, GPT-5.5 reaches 25.0% at the xhigh reasoning setting.
Models are also quickly improving, with the pass rates for the GPT-5 family increasing from 3.5% to 9.4% to 19.0% to 25.0% from 5.0 to 5.2 to 5.4 to 5.5.
Even the two strongest Pro settings remain far from saturated, with GPT-5.4 Pro and GPT-5.5 Pro leaving 62.1% and 60.2% of problems below 20% pass rate, respectively.
Pass rates also decline sharply with the length of the inferential chain. Problems with 3-4 decision points are solved at materially higher rates than those with 7+, and on the longest-chain problems the pass rate remains at 0% across every model evaluated. The gradient is steepest for the strongest models: weaker models sit near the floor regardless of chain length, while stronger models solve shorter chains reliably but break down as the number of required sequential correct inferences grows. This is consistent with the idea that errors compound through the chain; each additional fork is an independent opportunity to commit to a plausible-but-wrong choice, and the probability of threading all of them correctly falls off quickly.
Qualitative impressions on model improvements
It is useful to note the manner in which passing runs differ from those that fail and how well the pattern maps to what one observes in human scientists. In a manual review of selected GPT-5.4 and GPT-5.5 traces, we observed that both models are usually able to identify the presence of a data issue or a clue that a certain model is more appropriate than another. The main axis of improvement for stronger models arises downstream: stronger models are more likely to consistently allow these observations to change the actual analysis and therefore to steer its trajectory toward the correct answer. Relatedly, stronger models tend to perform significantly more exploratory data analysis and visualization, and are more likely to think through alternative hypotheses; weaker models are noticeably less "curious" and more single-minded.
Thus, the improvement from weaker to stronger models appears less like a simple increase in factual recall or execution capability and more like movement along a more researcher-like axis: better problem representation, better prioritization of anomalies, and better follow-through when a diagnostic should force a change in the analysis.
As a simple example, the target in one GeneBench problem is to compute polygenic scores for a set of individuals where the file with the set of weights to be applied includes a number of data inconsistencies.
One of these inconsistencies is that the same genetic indel variant is present in both the weights file as well as the file containing the genotypes to be scored, but has a slightly different representation in the two datasets (a common occurrence in real-world setups, as a given genetic variant can be represented in an arbitrary number of ways, and different tools output data with different formatting conventions).
For example, one of these indels is coded (in terms of chromosomal position, reference allele -> alternative allele) as chr22:30000000, ATTT -> A in the genotype manifest, while it is represented as chr22:30000001, TTTT -> T in the weights file. Both of these representations refer to the same underlying haplotype change (deletion of TTT) at the same genomic locus, but a naive join on position, ref, and alt would yield an empty intersection, and downstream scoring would omit the effect of this variant and therefore produce an incorrect result.
The correct approach thus requires recognizing this and normalizing the indel representations prior to merging the data. Both 5.4 and 5.5 consistently noticed something going on with these indels, but 5.4 was less able to identify these in applying the necessary corrective actions:
GPT-5.4: “Three weight entries (pos 30000001/30150001/30300001) had no matching genotype position and were dropped; therefore 3 genotype indels (rs_indel_1-3) received zero weight.”
GPT-5.5: “I standardized chr labels/case, matched exact alleles or strand complements for non-ambiguous SNPs, and rescued 5 indels whose representations differed because of normalization/left-right shifting...”
This qualitative pattern of stronger models being able to more consistently close this "notice-act" gap generally held across the traces examined.
Toward a future where verification is the bottleneck
My day-to-day job looks vastly different than it did a year ago; my responsibilities today resemble those in my previous role as a research team lead more than they resemble what an individual contributor would have been doing even two years ago. The difference is that instead of managing people, I manage and supervise agents.
Agents code much faster, work overnight, and enable analyses that previously would not have been practical because of the sheer amount of human effort they would have required. Yet the level of direction and micromanagement I find myself having to provide is still substantially higher than with humans (though this decreases somewhat with each model release), and we are clearly still a ways from agentic systems routinely displaying the level of judgment, intuition, and attentiveness that an experienced scientist might be expected to show. Reliability is especially important when stakes are high, and as long as a human has to perform this level of oversight, scientific research will remain bottlenecked by the human element.
GeneBench is an initial attempt to evaluate the capabilities of current models to perform such end-to-end quantitative scientific analyses, but it is not without its limitations. Real scientific problems involve a level of ambiguity, messiness, and shifting context that no current benchmark fully captures, and there is often not a single unambiguous "correct" answer to begin with.
Nevertheless, if the current pace of capability improvements continues, I anticipate that within the next few model releases, many of the current shortcomings will simply vanish through capability improvements in one axis or another, and that within a year or two, most industry and academic scientists will have shifted the primary load of such tasks entirely onto AI agents.
When that happens, the next bottleneck will be the same one that software engineers are currently dealing with: how to process, verify, and review the vast amounts of output that these agents will generate. The role of the average scientist will shift to a higher level of abstraction than data analysis: individual scientists will be able to explore multiple hypotheses in parallel using teams of agents and have confidence in their results, enabling exploration of larger sets of possible research directions that previously had to be ruthlessly trimmed and prioritized in the name of efficiency.